COMPANY NEWS
Patent in USA
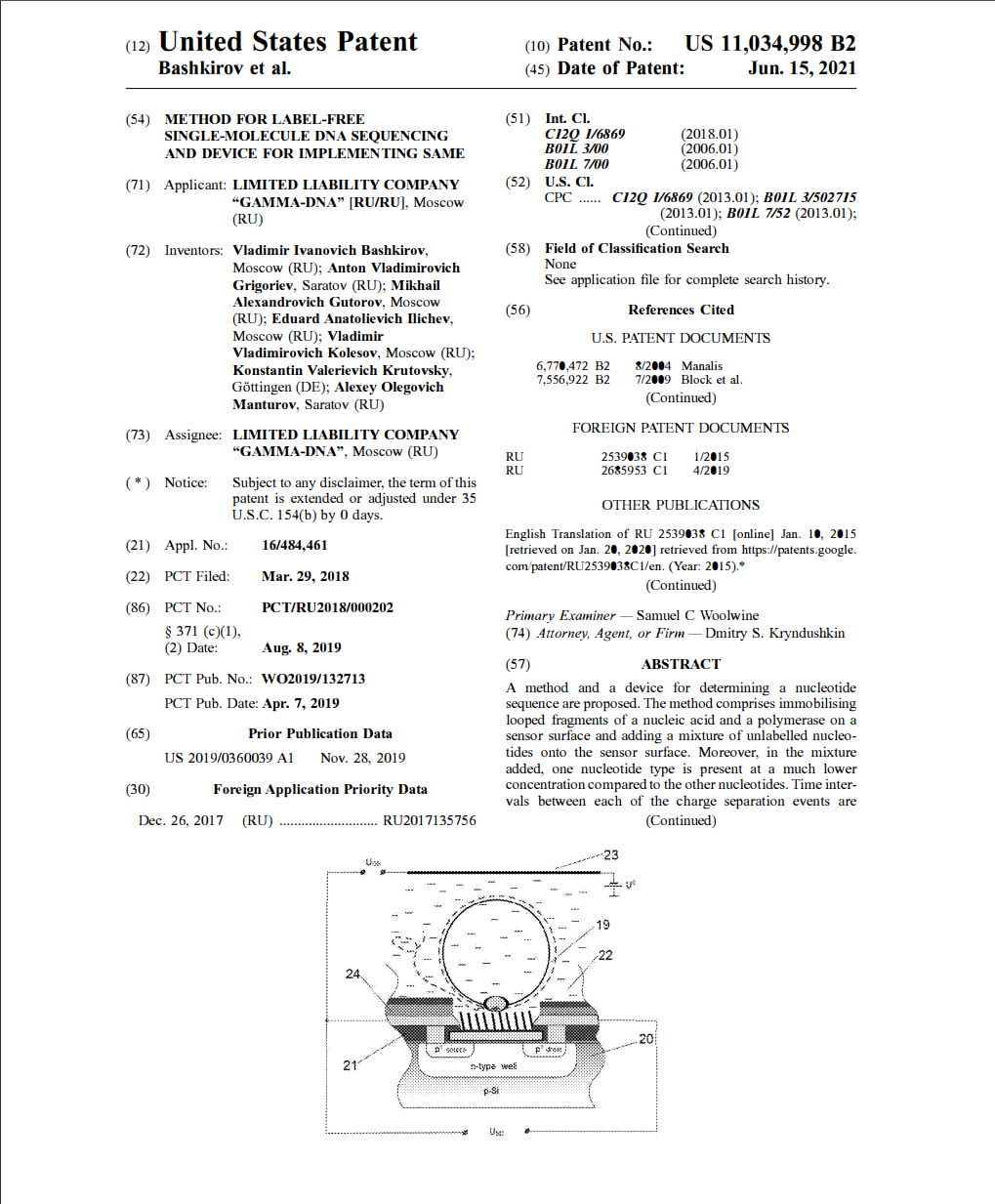

We are glad to announce that we were granted a patent for the "Method of unmarked single-molecule DNA sequencing and the device for its implementation" in USA!
In the patent the way and the device of determination of the nucleotide sequence of a molecule of nucleinic acid is offered.

Demonstration of a program that is designed to recover the nucleotide sequence using modelling data
At present, the debugging of the layout of the key nodes of the DNA sequencer necessary for conducting experiments on sequencing of custom oligonucleotide is being completed, the results of which will be formed and demonstrated by the program that we have developed for this task Today we provide
opportunity for site visitors to get acquainted with the work of this program using the example of synthesized data that emulates the sequencing data of a custom oligonucleotide using the layout of the key nodes of the DNA sequencer.
The program implements the original diffuse nucleic acid sequencing algorithm, which works with four source data files. The data for these files are generated by digitizing the sensor signal, which registers the facts of charge separation as a result of embedding (immobilized on the sensor surface) polymerase of each nucleotide during the polymerization reaction of a looped single-stranded oligonucleotide alternately in four different working solutions, respectively. Working solutions of nucleoside triphosphates are alternately fed to the sensor surface, each of the solutions having one type of nucleoside triphosphates, the molecules of which are concentrated to limit the rate of synthesis of the custom oligonucleotide. According to the sequencing algorithm, on average, the time intervals are longer before embedding the nucleotides of a species whose concentration was reduced against the normal concentration of the nucleotides of three other species in the working solution. Since, at the sample preparation stage, the name of the nucleotide species is known, its concentration is reduced for each working solution, the software tools for the key design of the DNA sequencer key nodes form time sequences for the sensor to record oligonucleotide synthesis results for each working solution in the form of a sequence of discrete time intervals containing logical units “ 1 "and logical zeros" 0 "; moreover, the logical unit "1" denotes the time interval that corresponds to the embedding of nucleotides, and the logical zero "0" denotes the time intervals when the reaction results were not recorded. The sequences of logical units “1” and logical zeros “0” thus obtained are recorded in four files corresponding to four working solutions. The program sequentially processes the data in four files, using the original algorithm sets the threshold value of the time interval for the time sequence of each file; with the threshold value all time intervals are compared between logical units “1” on the time sequence and sorted into long and short. Long time intervals correspond to the incorporation of known nucleotides, nucleotides of the species, the concentration of which has been lowered in the working solution
You can get acquainted with the work of the program by watching the video, which we specially recorded for this.

We are glad to announce that on February 11, 2019, we were granted a patent for the "Method of unmarked single-molecule DNA sequencing and the device for its implementation"!

We received a certificate of the computer program state registration №2018611229 called “The program of logic-mathematical 3D modeling of DNA polymerization reaction”.

FEBRUARY:
The Grant Committee of the Skolkovo Foundation approved a non-dilutive financing of “GAMMA-DNA LLC” according to the results of the expertise and a pitch of the project.

MARCH:
An application for an international patent for an invention (PCT) called “Method of Single-molecule Label-free DNA Sequencing and device for its realization” was filed.

APRIL-MAY:
There was an active interaction with investors (business angels and funds) and partners (Zethea, Moscow state University, NauTech, TFK Skolkovo, Skolkovo IP center), discussion of investment conditions and conditions of cooperation with partners.
JUNE:
Deal structuring with business angels, closing of the pre-seed round first stage.
Four business angels joined our project. They enriched the project with competencies in the field of artificial intelligence, nanotechnology and finance.
JULY:
The development within the pre-seed round was started according to the plan. We’ve purchased the equipment and prepared the laboratory; agreed technical specifications for polymerase and sensors with partners; concluded the agreements with partners.
We’ve also developed all technical requirements and methods, experimental design documentation for sensors, the microfluidic device, as well as for measuring stand and a layout of the key nodes of the DNA sequencer.
We took part in the industry-specific event – 2nd international conference "Bioinformatics: from algorithms to application" in St. Petersburg.
AUGUST:
We have opened a laboratory, staffed a team by biochemists, engineers, specialists in the field of information technology. Our mentors are actively involved in the work.
The preparation of technical requirements for the design of a prototype sensor cell matrix using standard TSMC technology was started.
SEPTEMBER:
Technological, business and legal expertise of the project by G&S I Fund LP (USA) for the further possible investment of the 2nd stage of the pre-seed round was carried out.
The development of the physical and mathematical model of the signal formation for sensors.
OCTOBER:
G&S I Fund LP (USA) took a positive decision to participate in the project and invest GAMMA-DNA LLC.
Charles Edward Shearer, General Partner, G&S I Fund LP (USA):
We have long been interested in the breakthrough venture projects in the field of medicine with a rich technological basis and high potential for commercial success. We were happy to make a positive decision about investing in the high-tech project, presented by GAMMA-DNA LLC. Our decision was based on the results of the deep due diligence, as well as the communication with the project team. We assessed the risks as acceptable for us at the pre-seed stage both due to the technological perspective of the project and its feasibility and due to the experience and professionalism of the project team: each of the technological directions and the direction of the business development are headed by experts with the deep knowledge and practical experience in their professional fields.
The uniqueness of the technology and its strategic potential to competitors are impressive. We intend to assist GAMMA-DNA LLC in building commercial and partnership relations and entering the markets of Asia and the USA, including licensing and promotion.
Nadya V. Matveyeva, Head of Business Development and Investor Relations, GAMMA-DNA LLC:
G&S I Fund LP is an important strategic partner for us – both in assisting in entering foreign markets and in the attraction of foreign investment for the subsequent rounds. The US market is monopolized by the leader (Illumina) and initially was not in the list of our strategical potential foreign markets. But with the assistance of the G&S I Fund LP team, the issue of representation in the US may well be considered.
Our key task today is to ensure for all the shareholders of the company the optimal balance between the volume of new investments attracted and the speed of R&D and business development in order to reduce the time to market.
Also, in October we’ve developed technological regulations for the manufacturing of two types of sensors and we’ve developed the analog-to-digital circuit and signal processing topology for charge-sensitive matrix cell in accordance to TSMC technology in October.
Participation in the 7th Moscow International Forum «Open Innovations» at the SKOLKOVO innovation center. In the competition of projects "We are from NTI" project "GAMMA-DNA" took 1st place in HealthNet unanimously! Find more details – here.
NOVEMBER:
The first batch of polymerase that was manufactured by Zethea according to the technical specifications of GAMMA-DNA was received. The polymerase corresponds to the planned characteristics.
We received prototypes of two variants of the sensor manufactured by MSU according to the technical specifications of GAMMA-DNA. Sensors parameters correspond to the planned.
The development of software to control the layout blocks of the key nodes of the DNA sequencer was started. Also, we’ve started started the development of software for the formation and demonstration of the DNA fragment sequencing results by the layout of the key nodes of the DNA sequencer.
DECEMBER:
The measuring stand was manufactured and tested. It is designed to test for the serviceability of two variants of sensors and for measuring their electrical parameters. The measuring stand is necessary to select the best sensor from two variants of sensors for use it in subsequent experiments with biological material.
We’ve also developed the concept of experiments to demonstrate the results of the ordered oligonucleotide sequencing. The oligonucleotide is already been bought and is stored in the freezer.
The know-how of the method of DNA samples preparation for use in a single-molecule DNA sequencing was registered. Also we’ve registered the know-how of the unique algorithm of a single-molecule DNA sequencing.
Michael Gutorov, CEO, Head of Physics and Electronics, GAMMA-DNA LLC
From the point of our R&D, 2018 made it possible to make a significant progress in the work and to make closer the day of the demonstration of a sequence of the ordered oligonucleotide on the layout of the key nodes of our DNA sequencer (prototype).
Planned R&D and business development tasks were completed. The patent application for international status (PCT) for the invention “Method of Single-molecule Label-free DNA Sequencing and device for its realization” was filed ahead of a schedule.
Each department leader has formed a team of specialists who have successfully passed the trial period. Today, the staff of employees is 17, also our scientific consultant and 2 mentors work with us.
The laboratory is equipped with all the necessary equipment.
With such results and resources, we’ve confidently stepped into 2019!
GAMMA-DNA LLC won the 1st place in HealthNet at Open Innovations 2018!

at the entrance

at the exit

Polymerase development: we’ve started - in partnership with ZETHEA